NMPDR Search
The NMPDR search facility provides many different ways to look for genes, genomes, or sequences in the NMPDR database. The following searches are currently supported.
| DrugSearch | Show in silico docking results for select PDBs. |
| FidSearch | Display genes from one or more genomes filtered by subsystem or keywords. |
| OpSearch | Display genes of a single genome organized into operons. |
| SigGenes | Display genes that are common to a set of genomes, or that differentiate between two sets of genomes. |
| SubSearch | Search for genes in a given subsystem or subsystem class. |
| ToolSearch | Search for genes or DNA using BLAST or pattern-matching. |
| WordSearch | Display genes that match certain keywords. |
The results of a search will generally start with data for the NMPDR core organisms. All NMPDR searches allow you to download results in tab-delimited and XML formats. NMPDR searches that return a list of genes also allow you to download the genes in FASTA format.
Most NMPDR searches use a powerful Genome Control that allows you to choose one or more genomes according to various criteria. Some also use a search-engine style Keyword Box. The Keyword Box can also be found at the top of many of the NMPDR pages.
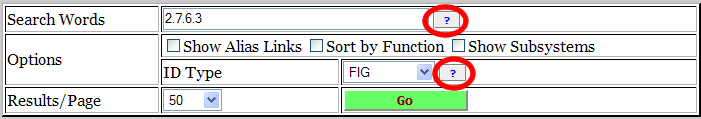
If you have a question about something in an NMPDR search form, mouse over the a question mark to get a hint. Click on the blue question mark to pop up the Wiki page on the subject. The blue question marks have been circled in the sample search form below.