SEED Viewer Manual/Editing Capabilities/ChromosomalClusters
Editing Capabilities - Chromosomal Clusters
This page can be accessed from a Chromosomal Clusters view, using the button annotate clusters (only visible if you have editing rights for the genome).
Overview Table
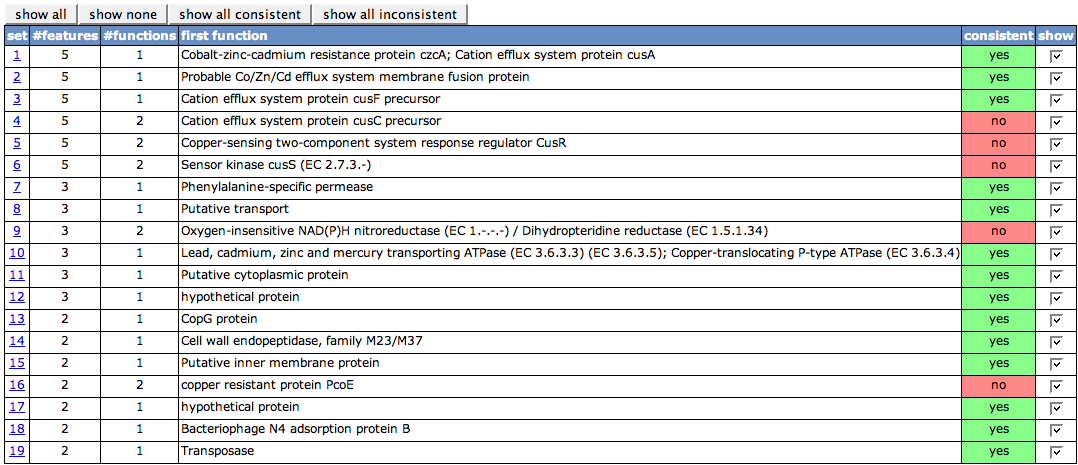
The first table on the page lists all sets of protein visible in the Chromosomal Clusters view. The set numbers in the first column are the set numbers from the view. The second column (#features) shows the number of features in the set. #functions depicts how many different functions the features in the set have. One of the functions is displayed in the column first function. Whenever there is more than one function, the cell in the next column will be painted in red, as it is not consistent, else green. With the check boxes in the last column, you can turn on and off the details tables for each of the sets.
Above the table, there are four buttons to turn on/off the details tables. show all will remove previous filtering and show you all details tables again. show none will remove all details tables, so that you can afterwards select single ones you'd like to see. show consistenct and show inconsistent is related to the consistent column in the table. It will display the details tables with a consistent or an inconsistent set, respectively.
As a default, for each set you will get a details table in the following. These list all the features in the set. The set number is displayed in the first column. Organism is the genome the feature stems from. If there are more than one occurrence of features in the set for one genome, they are numbered in the Occ column. If there is a UniProt alias for the feature, the ID and the function of this alias will be shown in the next two columns.